CRISPR-based gene and genome modification technology is revolutionizing targeted approaches in plant genetics. Combined with single-cell analytics, it enables the generation and testing of exciting hypotheses on cell-type-specific processes (Pfeiffer et al., 2022).
Developmentally controlled programmed cell death (dPCD) exemplifies a cell-type-specific process, as it typically occurs in a single cell within a specific tissue. Therefore, both single-cell analyses and cell-type-specific modulation of dPCD are essential for our research.
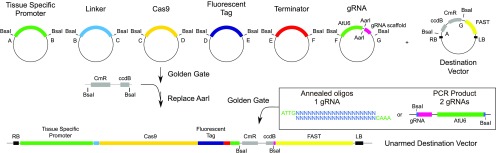
In collaboration with Thomas Jacobs and Tom Beeckman (VIB-UGent), we have been developing CRISPR technology for precise gene modifications in specific cell types.
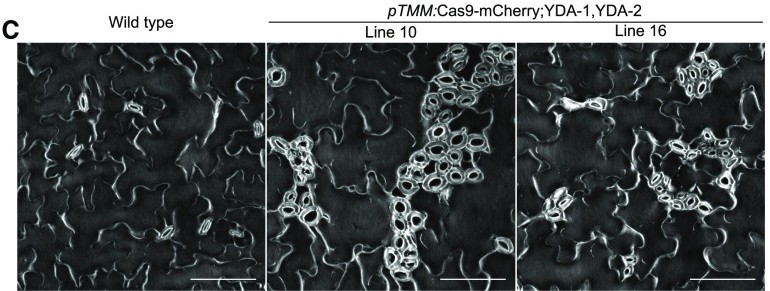
Our CRISPR-based Tissue-Specific Knock-Out (CRISPR-TSKO) approach has proven highly effective in multiple tissues and cell types, including the root cap, stomatal lineage, and lateral roots (Decaestecker et al., 2019). Notably, the system is also highly efficient with gRNA multiplexing, enabling the simultaneous knock-out of up to six genes (Bollier et al., 2021).
Since CRISPR-TSKO generates mosaic genomic edits, thorough evaluation is crucial. Cell sorting and DNA sequencing are essential for assessing outcomes, with guidelines available in this Methods in Molecular Biology book chapter (Decaestecker et al., 2022).
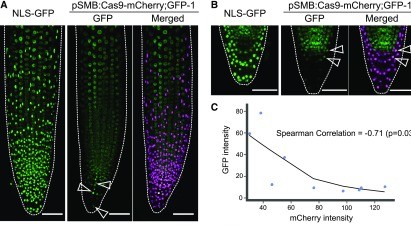
In addition to CRISPR-TSKO, we are also collaborating with Thomas Jacobs (VIB-UGent) in the development of multiplex combinatorial CRISPR screens, targeting sets of dozens or hundreds of genes (e.g. gene families, functional groups of genes, ...) in a randomized combinatorial fashion. This approach enables us to screen for higher-order mutant phentoypes, combining the advantages of forward and reverse genetics (lead scientist Marie L. Pfeiffer, unpublished).