Root Cap Turnover as Powerful Model System to Study Developmental Programmed Cell Death
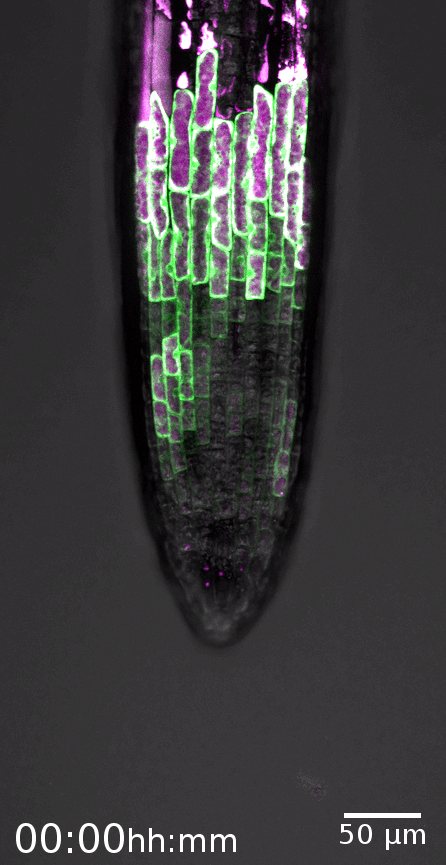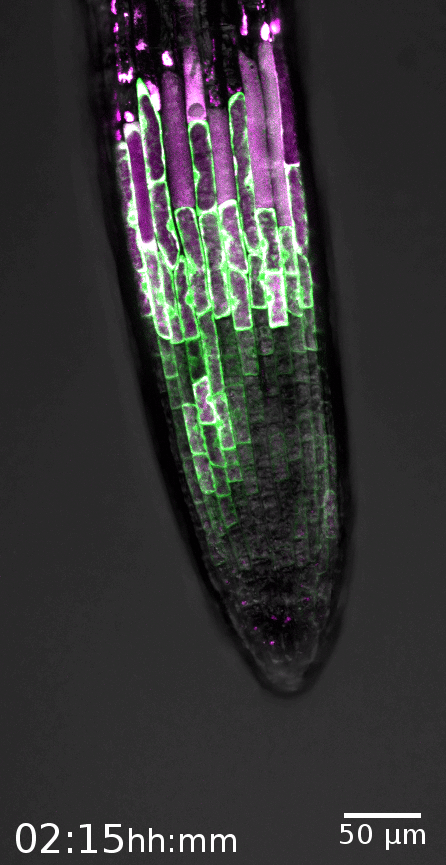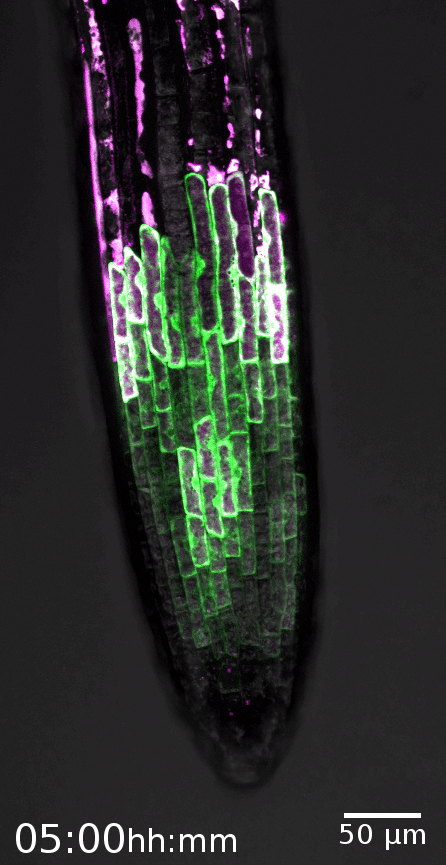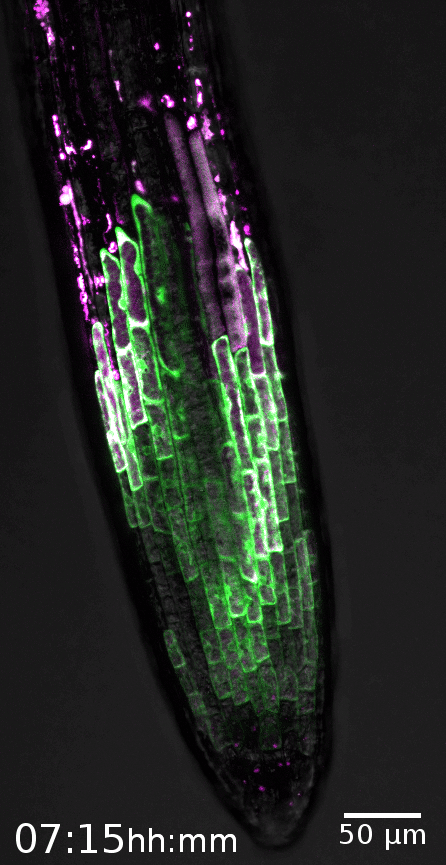
The root cap is the signature model system of the Nowack lab. This specialized organ covers the growing root tips of nearly all vascular plant species, serving crucial functions such as gravity perception, lubrication of the root growth canal, and communication with the rhizosphere biome (Vahldick et al. 2024). Typically, it consists of multiple layers covering the root meristem, formed by two distinct cell types: columella root cap cells at the very tip and lateral root cap cells at the flanks (Kumpf and Nowack, 2015).
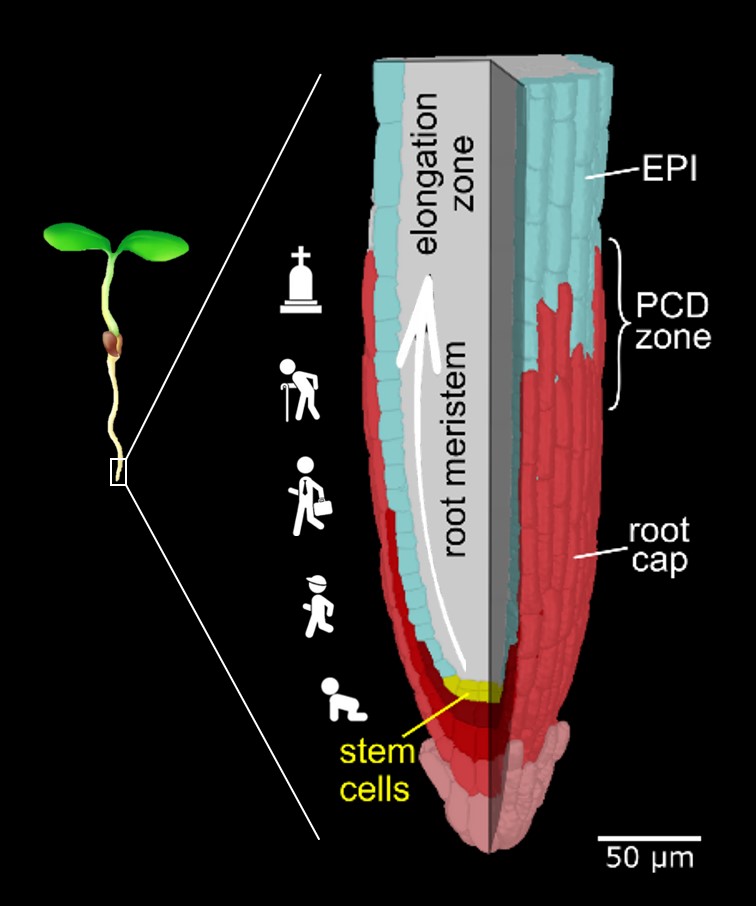
By identifying dPCD signature genes and analyzing their expression patterns in Arabidopsis thaliana (Olvera-Carrillo et al., 2015), we established that root cap turnover is regulated by a balance between cell generation by specific stem cells and degeneration via dPCD (Fendrych et al., 2014). In collaboration with Tom Beeckman (VIB Ghent), we proposed that dPCD in lateral root cap cells releases pulses of auxin into the root, contributing to the spatial patterning of lateral root formation (Xuan et al., 2016). Further, with Christiane Nawrath (UNIL), we identified a novel cuticle-like layer covering the outermost embryonic root cap layer and lateral root caps, facilitating emergence and protecting young root organs (Berhin et al., 2019).
Building on these discoveries, we established root cap turnover as a leading model for dPCD research. We identified an extensive gene regulatory network that coordinates root cap differentiation, maturation, and preparation for dPCD execution (Fendrych et al., 2014, Huysmans et al., 2018, Feng et al., 2023).
The downstream output of these regulatory networks includes the dPCD signature genes. However, their precise roles in cell death execution remain largely unknown, as genetic manipulation of these genes does not produce obvious cell death phenotypes. A notable exception is the nuclease BFN1: loss of BFN1 function delays post-mortem clearance, confirming its role as an integral component of genetically encoded dPCD (Fendrych et al., 2014).
Live-Cell Imaging and the Cell Biology of dPCD Execution
The root cap's accessibility to live-cell imaging and pharmacological approaches has allowed us to investigate dPCD execution with unprecedented spatial and temporal resolution. We described a highly reproducible decompartmentalization process that enables rapid dPCD execution within minutes. Early events include leakage of mitochondrial matrix, nuclear contents, and ER lumina into the cytosol. However, classic dPCD hallmarks—such as plasma membrane permeabilization and vacuole collapse—occur much later in the process (Wang et al., 2024).
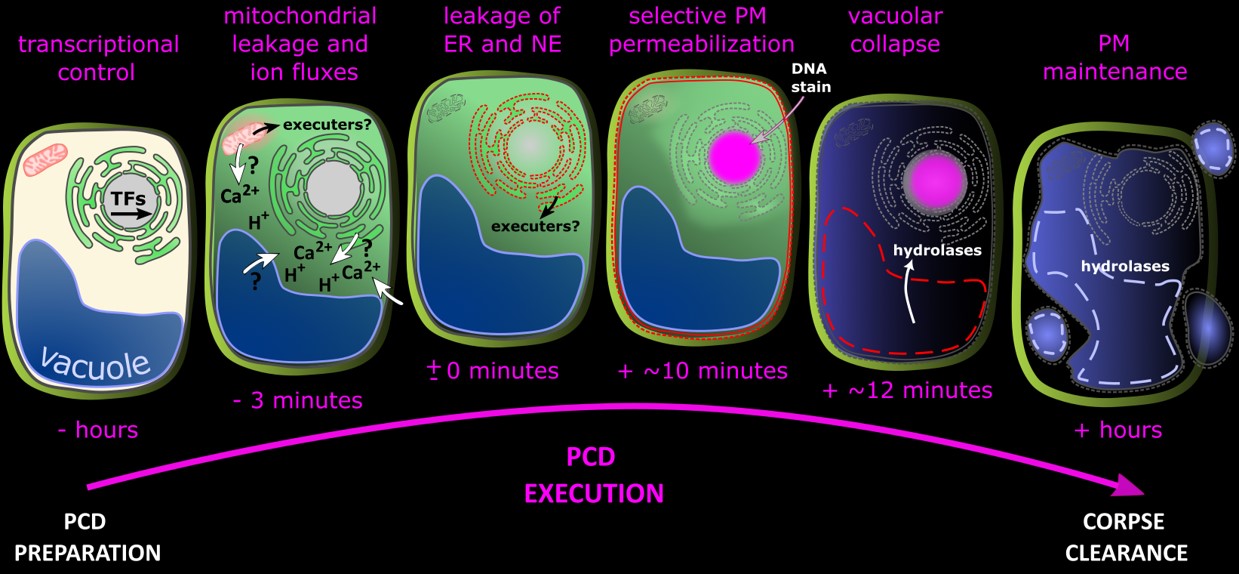
These findings pave the way for formulating and testing new hypotheses on the molecular executioners of plant dPCD.
Autophagy and dPCD
In collaboration with Yasin Dagdas (GMI Vienna, now COS Heidelberg), we established that autophagy is upregulated prior to root cap dPCD execution. However, autophagy plays divergent roles in different root cap cell types. In some, autophagy-deficient mutants exhibit delayed cell death and post-mortem corpse clearance, while in lateral root cap cells, dPCD proceeds independently of autophagy (Feng et al., 2022).
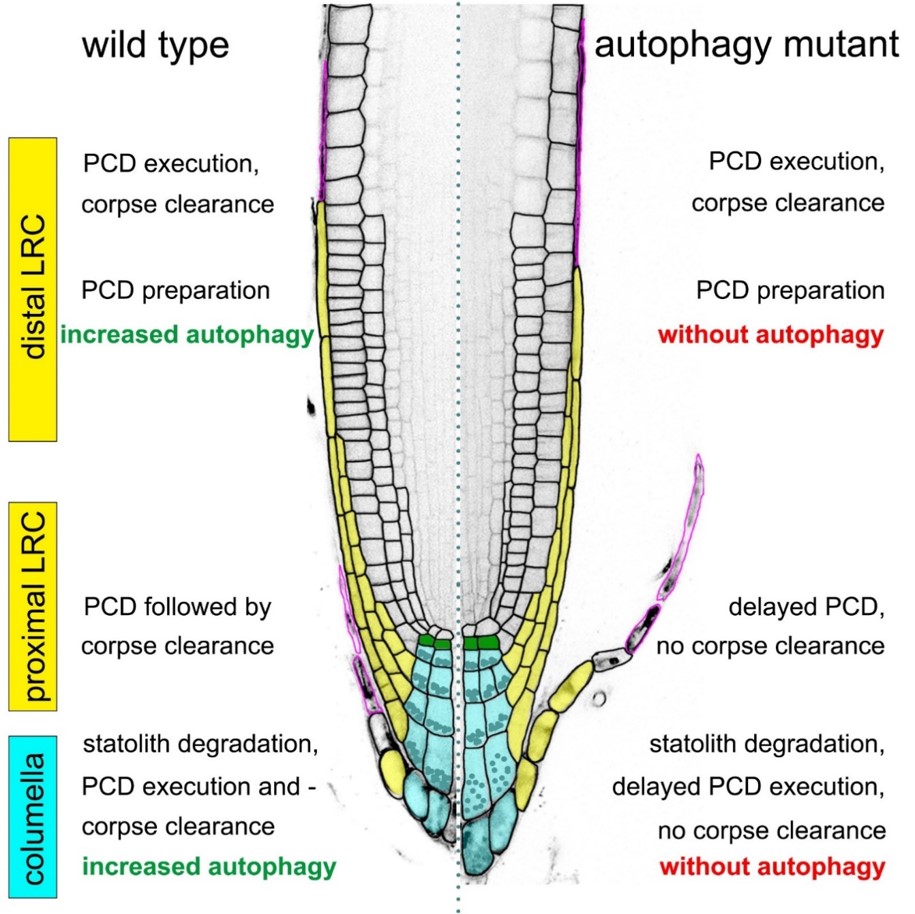
Root Cap dPCD and the Microbiome
Beyond its developmental roles, root cap dPCD also influences interactions with the root microbiome. In collaboration with Alga Zuccaro (University of Cologne), we demonstrated that tight regulation of post-mortem corpse clearance is crucial for limiting microbial colonization of the root and rhizosphere in Arabidopsis (Charura et al., 2024). Future research in this area will further explore these interactions.
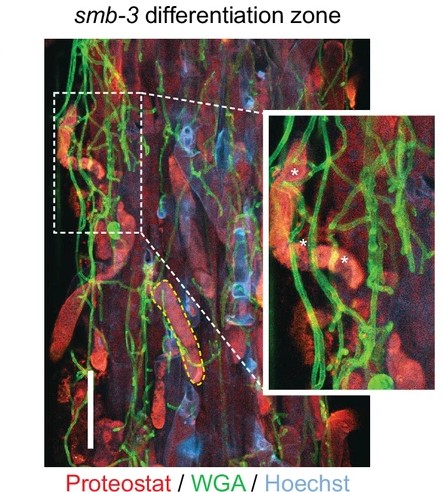
References
Kumpf, R. P. & Nowack, M. K. The root cap: A short story of life and death. J. Exp. Bot. 66, 5651–5662 (2015). doi:10.1093/jxb/erv295
Olvera-Carrillo, Y. et al. A conserved core of PCD indicator genes discriminates developmentally and environmentally induced programmed cell death in plants. Plant Physiol. pp.00769.2015 (2015). DOI: 10.1104/pp.15.00769
Fendrych, M. et al. Programmed cell death controlled by ANAC033/SOMBRERO determines root cap organ size in arabidopsis. Curr. Biol. 24, 931–940 (2014). DOI: 10.1016/j.cub.2014.03.025
Xuan, W. et al. Cyclic programmed cell death stimulates hormone signaling and root development in Arabidopsis. Science. 351, 384–387 (2016). DOI: 10.1126/science.aad2776
Berhin, A. et al. The Root Cap Cuticle: A Cell Wall Structure for Seedling Establishment and Lateral Root Formation. Cell 176, 1367-1378.e8 (2019). DOI: 10.1016/j.cell.2019.01.005
Huysmans, M. et al. NAC transcription factors ANAC087 and ANAC046 control distinct aspects of programmed cell death in the arabidopsis columella and lateral root cap. Plant Cell 30, 2197–2213 (2018). OI: 10.1105/tpc.18.00293
Feng, Q. et al. Repressive ZINC FINGER OF ARABIDOPSIS THALIANA proteins promote programmed cell death in the Arabidopsis columella root cap. Plant Physiol. 192, 1151–1167 (2023). DOI: 10.1093/plphys/kiad130
Wang, J. et al. A developmentally controlled cellular decompartmentalization process executes programmed cell death in the Arabidopsis root cap. Plant Cell 36, 941–962 (2024). DOI: 10.1093/plcell/koad308
Feng, Q., De Rycke, R., Dagdas, Y. & Nowack, M. K. Autophagy promotes programmed cell death and corpse clearance in specific cell types of the Arabidopsis root cap. Curr. Biol.32, 2110-2119.e3 (2022). DOI: 10.1016/j.cub.2022.03.053
Charura, N. M. et al. Root cap cell corpse clearance limits microbial colonization in Arabidopsis thaliana • Cologne Excellence Cluster for Cellular Stress Responses in Aging-Associated Diseases Key findings in bullet points. eLife (2024). https://doi.org/10.7554/eLife.96266.3
Collaborators
https://www.unil.ch/dbmv/en/home/menuinst/research/recherche/dr-christiane-nawrath-mer.html
https://www.oeaw.ac.at/gmi/research/research-groups/yasin-dagdas/profile
https://ag-zuccaro.botanik.uni-koeln.de/